Gene editing technology is the core technology of Biocytogen. Our independent research and development efforts have significantly enhanced the efficiency and success rate of gene editing, overcoming limitations in DNA length for gene modification. These original technologies have enabled us to develop a range of core animal/cell models that aim to address key challenges in drug development and improve clinical translation success rates. By optimizing our underlying logic, we have created a brand-new model for antibody drug development.

Biocytogen's gene editing platform was established in 2009 to engineer animal and cell models for clients.
These genetically engineered models are crucial not only for basic scientific research but also for solving key problems in the process of antibody drug research and development. For instance, Biocytogen's fully human antibody RenMab mice can directly obtain fully human antibody sequences screened by the immune system. Additionally, their HCAb RenNano mouse is soon to be used to develop fully human nanobodies. Furthermore, Biocytogen's gene editing models have a wide range of functions such as improving immune response to antigens by modifying certain genes or inducing disease models through genetic modification.
After years of independent technology research and development, Biocytogen has established three major gene-editing technology systems: (1) ESC/HR technology based on mouse embryonic stem cells that allow multiple rounds of genetic modification on the same chromosome; (2) EGE technology that increases CRISPR/Cas9-mediated gene knock-in efficiency by nearly 20-fold; (3) SUPCE technology that enables super-large gene segment humanization with up to three mouse embryonic stem cell manipulations without being limited by DNA segment length. By combining these three systems, Biocytogen has overcome common problems in gene editing technology while achieving high efficiency, large-scale multi-modification, and super-large gene humanization. Precise genome editing at almost any length and site can now be achieved thanks to their innovative approach.
ESC/HR
ESC/HR technology is a powerful tool for studying gene function and disease mechanisms in mice. By introducing specific genetic modifications into embryonic stem cells, researchers can create mouse models that mimic human diseases or test the effects of potential therapies. The use of homologous recombination ensures precise targeting of the desired genomic locus, minimizing off-target effects and increasing the efficiency of gene editing. The C57BL/6 background stem cell line used by Biocytogen is a commonly used strain in biomedical research due to its well-characterized genetics and susceptibility to many diseases. The ability to maintain totipotency after multiple passages allows for efficient production of genetically modified mice without compromising their health or viability. Overall, ESC/HR technology has revolutionized the field of mouse genetics and continues to be an essential tool for understanding complex biological processes and developing new treatments for human diseases.

EGE
The Extreme Genome Editing system (EGE) is a revolutionary gene editing technology that builds upon the CRISPR/Cas9 platform. Through independent research and optimization, EGE has increased the efficiency of gene knock-in by nearly 20-fold. While CRISPR/Cas9 simplifies gene editing operations and enhances homologous recombination probability, it remains inefficient and costly for large-scale model preparation. The EGE system streamlines DNA sequence editing at almost any genomic site, making it faster and easier than ever before. This heightened efficiency in gene editing offers numerous benefits such as increasing success rates, reducing costs, shortening cycles for model animal preparation - all of which are advantageous to industrial and commercial applications on a larger scale. Additionally, EGE technology is expected to improve the efficiency of modifying difficult-to-reach genomic sites thereby expanding its potential impact even further.
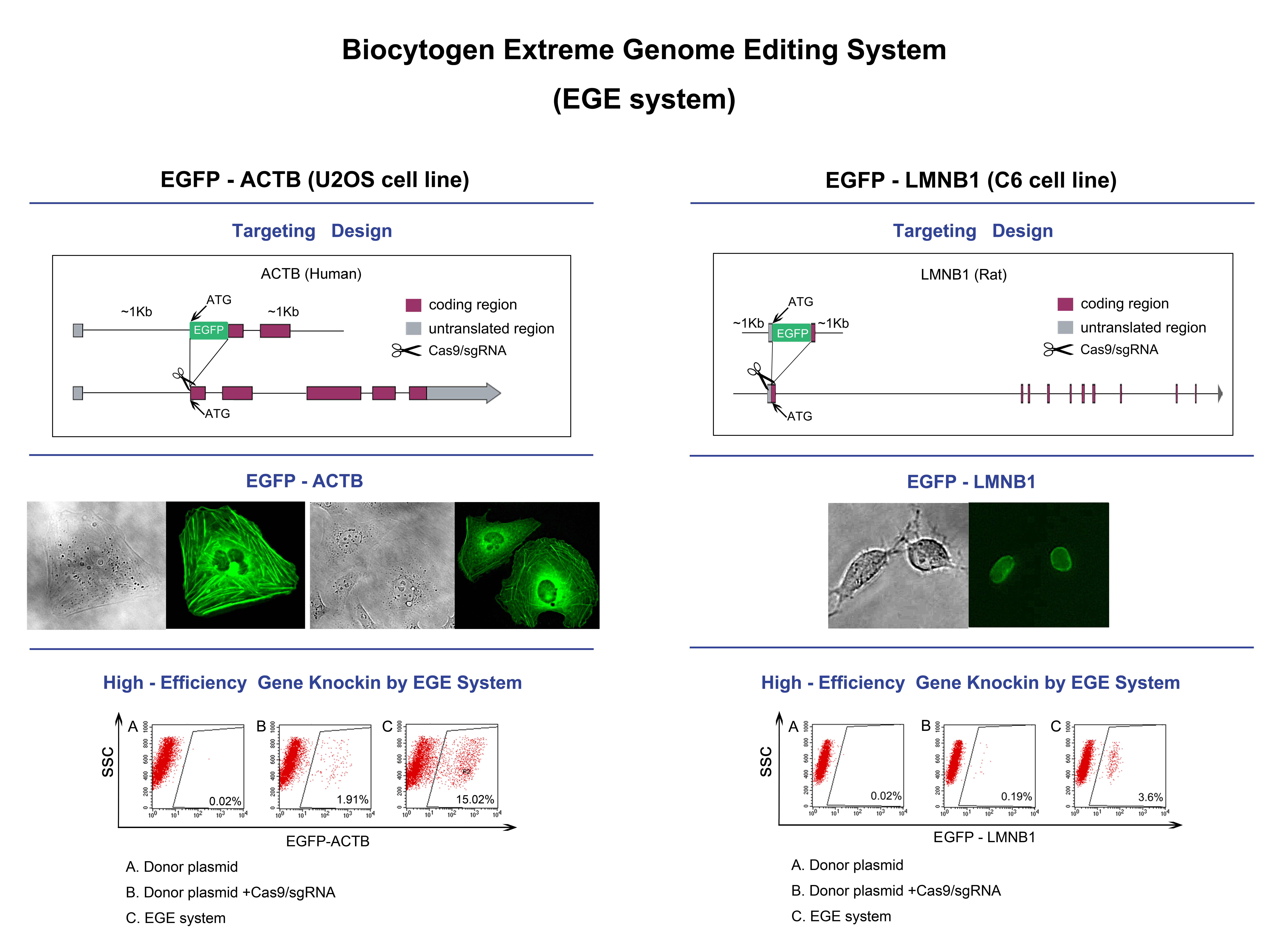
Figure 1. EGE technology greatly increases the efficiency of CRISPR/Cas9-based gene knock-in.
In the first model (left), human osteosarcoma cells are used. The design protocol is to knock in green fluorescent protein EGFP behind the initiation codon ATG of human cytoskeleton protein β-Actin gene ACTB. Cells successfully knocked in express the EGFP-fused β-Actin protein. U2OS cell transfection expresses Cas9 nuclease and sgRNA plasmid targeting ACTB, as well as EGFP donor plasmid containing homologous arms about 1 kb in length on both sides. After transfection, flow cytometry analysis demonstrates that the efficiency of basic Cas9/sgRNA-mediated knock-in is 1.91%, while the EGE system increases the knock-in efficiency to 15.02%. In the second model (right), the rat glioma C6 cell and the rat Lmnb1 gene encoding nuclear membrane protein are selected as targets. In this model, the EGE system increases the knock-in efficiency from 0.19% to 3.6%.
-
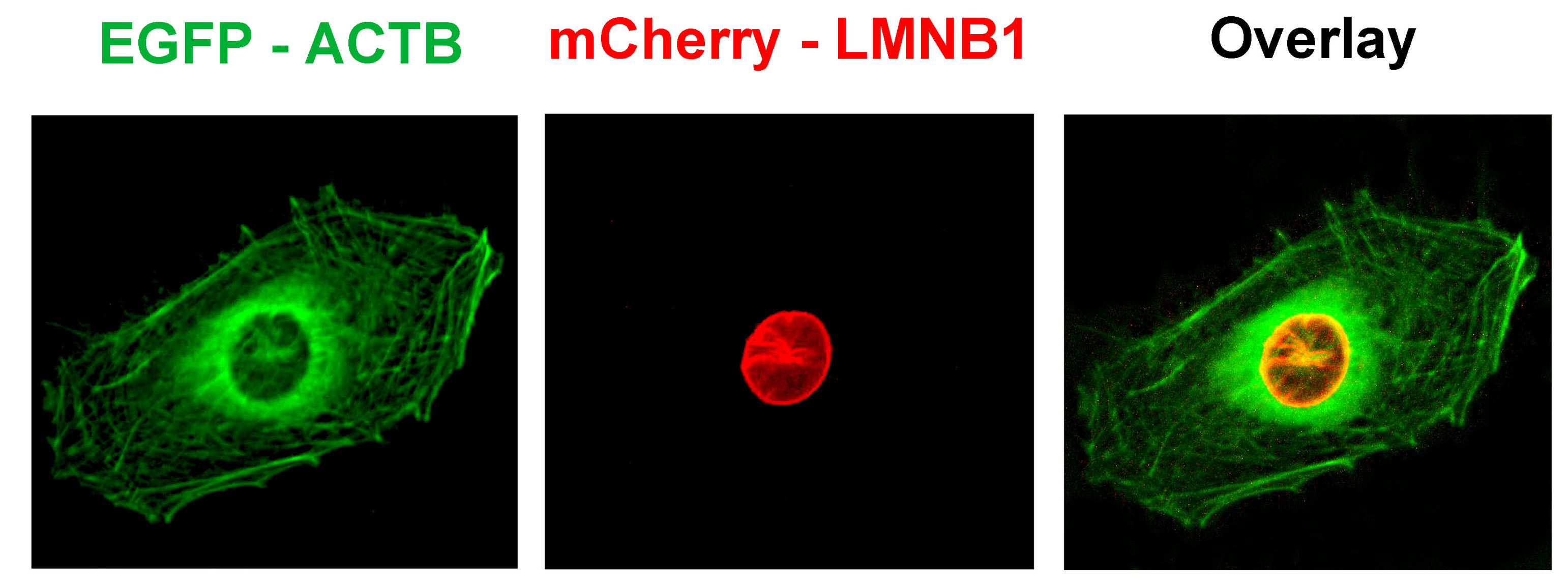
Figure 2. EGE technology allows two genes to be marked simultaneously in a cell.
-
In U2OS cells, co-transfection respectively targets ACTB, Cas9/sgRNA plasmid of LMNB1 gene, green fluorescent protein GFP donor plasmid marked with ACTB, and red fluorescence protein mCherry donor plasmid marked with LMNB1 gene. After transfection, the red nuclear membrane protein and the green cytoskeletal protein can be observed under fluorescence microscope.
SUPCE
SUPCE (Size-Unlimited and Precise Chromosome Engineering system) is a revolutionary genome-editing technology that overcomes the limitations of previous methods. Unlike plasmids or bacterial artificial chromosomes (BAC), SUPCE can replace large segments of DNA without requiring multiple rounds of modification in mouse embryonic stem cells. This eliminates the risk of losing totipotency due to long-term manipulation, which has been a major obstacle in obtaining adult mice using traditional techniques.
Biocytogen has successfully used SUPCE to develop RenMab, a fully human antibody mouse, as demonstrated by FISH test results shown in Figure 2. In addition to antibodies, SUPCE can also be applied to other super-large gene clusters such as T-Cell Receptor (TCR), Major Histocompatibility Complex (MHC), and NK Cluster (NKC). To support high-volume gene editing projects using these methodologies, we have established an automated workflow equipped with several 96-channel nucleic extractors for genomic DNA isolation and plasmid preparation. Our platform now enables the development of more than 1,500 models per year with rapid turnaround times and stable results.
-
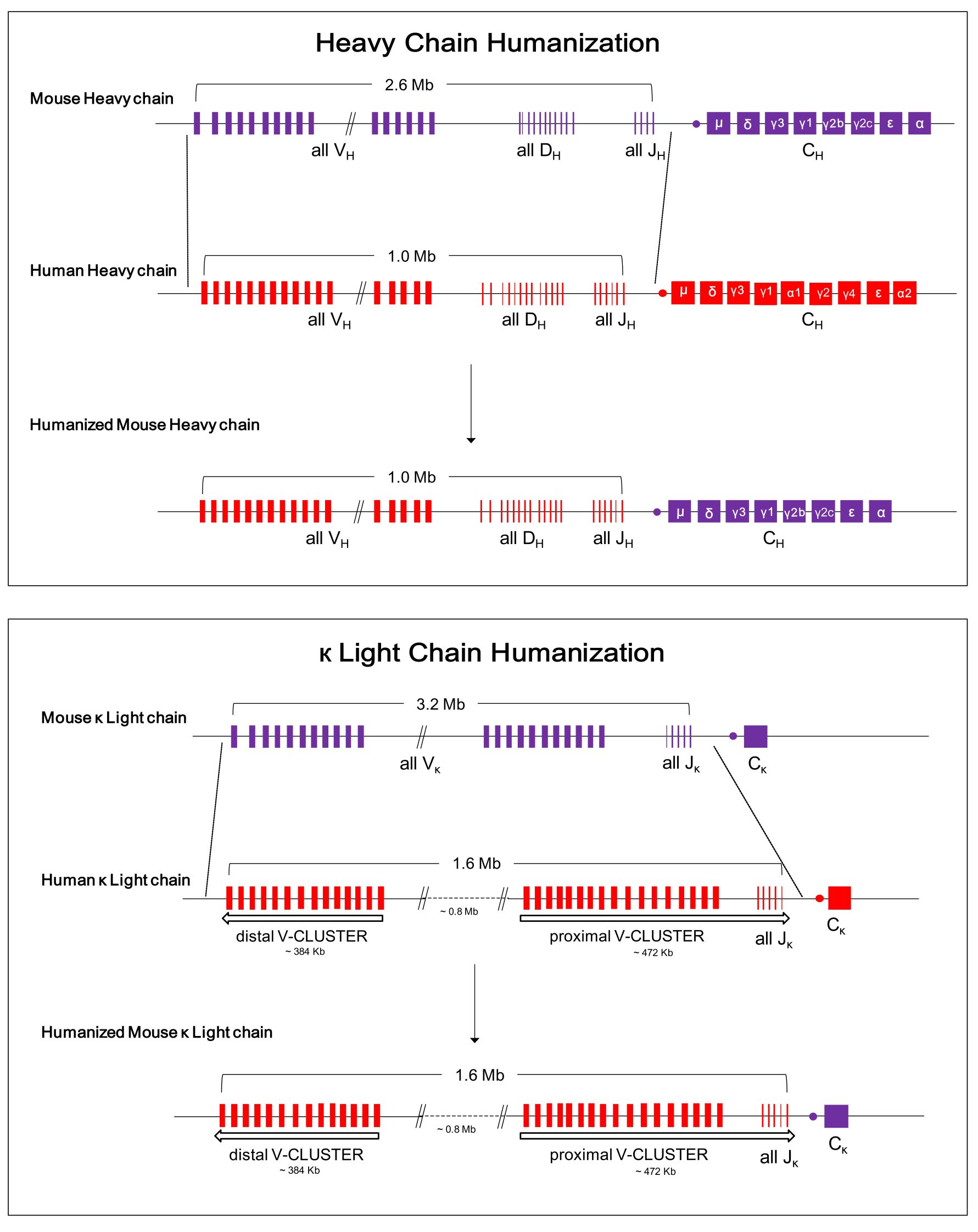
Figure 1.Schematic diagram of humanization design of antibody gene in RenMab mouse.
-
In RenMab mouse, the variable region of 2.6 Mb mouse heavy chain was replaced with the variable region sequence of 1.0 Mb human heavy chain and the variable region of 3.2 Mb mouse κ-light chain was replaced with the variable region sequence of 1.6 Mb human light chain at one time.
-
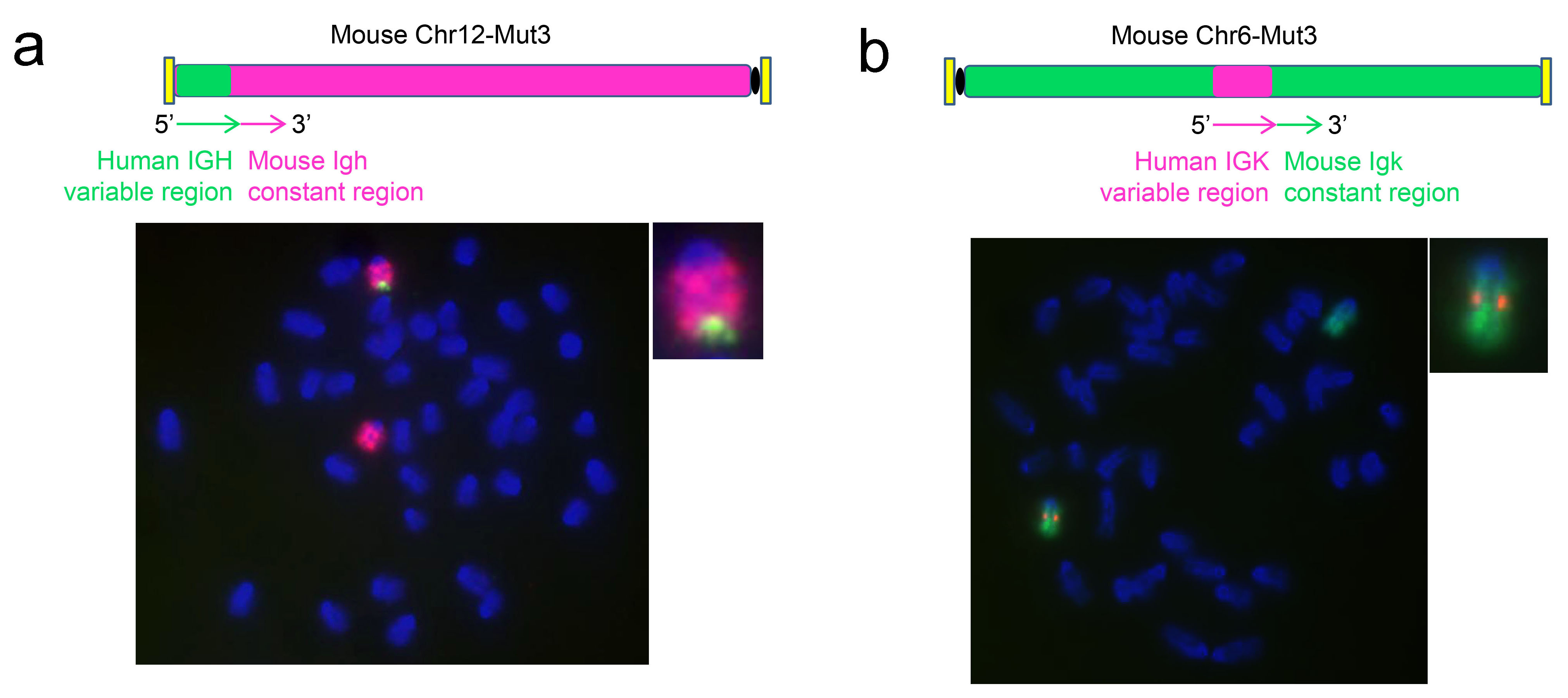
Figure 2.FISH test results of humanized replacement of variable region of antibody gene.
-
The variable regions of mouse heavy chain and κ-light chain were replaced with the human sequences in situ in mouse embryonic stem cells using SUPCE technology. A. With the variable region sequence probe (green) of human heavy chain, the chromosome #12 of mouse Igh was stained with the probe (red) to test the heavy chain humanized cell cloning. With the variable region sequence probe (red) of human κ-light chain, the chromosome #6 of mouse Igk was stained with the probe (green) to test the κ-light chain humanized cell cloning.
To support the high volume of gene editing projects using the above methodologies, we have established an automated, high-throughput workflow to ensure rapid, stable and large-scale model preparation. Our facility is equipped with several 96-channel nucleic extractors for the isolation of mouse/cell genomic DNA and plasmid DNA. The automated pipetting workstation accommodates the large volume of genotype testing performed. Our gene editing platform now enables the development of more than 1,500 models per year.